10 KiB
DICOM MCP Server
A Model Context Protocol (MCP) server for analyzing DICOM medical imaging files, specifically designed for QA workflows in body composition analysis and liver imaging.
Who is this for?
This server is designed primarily for Claude Desktop users — radiographers, QA analysts, and other non-technical team members who need to inspect and validate DICOM files but don't write code. Claude Desktop has no shell or filesystem access, so the MCP server is the only way to give it DICOM analysis capabilities.
Claude Code users (developers, engineers) generally don't need this. Claude Code can run Python and use pydicom directly, offering more flexibility than the predefined tools here. That said, the MCP can still be useful in Claude Code as a convenience layer if you want consistent, structured QA outputs without writing ad-hoc scripts each session.
What is this for?
This MCP server provides a plain-language API for interacting with DICOM data. It doesn't eliminate the need for Python/pydicom knowledge, but it significantly reduces the cognitive load for many common tasks.
What it does well:
- Makes common QA tasks instant (Dixon detection, header comparison, protocol validation)
- Removes boilerplate — no more writing the same pydicom scripts repeatedly
- Natural language interface — "find Dixon sequences" vs
pydicom.dcmread()loops - Pixel-level analysis — render images, measure signal statistics, compute SNR
- Consistent output formats across all operations
- Configurable PII filtering to redact patient tags when required
What it doesn't (and shouldn't) try to do:
- Replace pydicom for custom/novel analyses
- Handle every edge case in DICOM
- Be a full DICOM viewer/editor
It's the perfect "80/20" tool — handles 80% of routine QA tasks with 20% of the effort. For the remaining 20% of complex cases, users still have full Python/pydicom access.
Tools
The DICOM MCP server provides 17 tools across six categories:
Directory & File Discovery
| Tool | Description |
|---|---|
dicom_list_files |
List DICOM files with metadata filtering and optional count_only mode |
dicom_summarize_directory |
High-level overview with unique tag values and file counts |
dicom_find_dixon_series |
Locate and classify Dixon (chemical shift) sequences |
dicom_search |
Find files matching filter criteria (text, numeric, presence operators) |
Metadata & Validation
| Tool | Description |
|---|---|
dicom_get_metadata |
Extract header information organised by tag groups |
dicom_compare_headers |
Side-by-side comparison of up to 10 files |
dicom_validate_sequence |
Check acquisition parameters against expected values |
dicom_analyze_series |
Comprehensive series consistency and completeness check |
dicom_query |
Query arbitrary tags across all files with optional grouping |
Philips Private Tags
| Tool | Description |
|---|---|
dicom_query_philips_private |
Query Philips private DICOM tags by DD number and element offset |
Pixel Analysis
| Tool | Description |
|---|---|
dicom_read_pixels |
Pixel statistics (min, max, mean, std, percentiles, histogram) with optional ROI |
dicom_compute_snr |
Signal-to-noise ratio from signal and noise ROIs |
dicom_render_image |
Render DICOM to PNG with configurable windowing and ROI overlays |
Structure & Comparison
| Tool | Description |
|---|---|
dicom_dump_tree |
Full hierarchical dump of DICOM structure including nested sequences |
dicom_compare_uids |
Compare UID sets between two directories to find shared, missing, or extra UIDs |
Segmentation & T1 Mapping
| Tool | Description |
|---|---|
dicom_verify_segmentations |
Validate that segmentation files correctly reference their source images |
dicom_analyze_ti |
Extract and validate inversion times from MOLLI/NOLLI T1 mapping sequences across vendors |
For detailed usage examples, parameter reference, and QA workflows, see docs/USAGE.md. For a plain-English summary of what each tool does, see docs/CAPABILITIES.md. For behavioural constraints and regulatory context, see docs/GUIDELINES.md.
Installation
See INSTALL.md for complete installation instructions covering:
- Standalone package — No Python required, ideal for non-developers
- Claude Desktop — Poetry-based developer setup with GUI configuration
- Claude Code — Adding the MCP server to your Claude Code environment
- PII filtering — How to enable patient data redaction for team deployments
Quick Start (Developer)
# Install dependencies
poetry install --with dev
# Run tests to verify
poetry run pytest -v --tb=short
# Start the server
poetry run python -m dicom_mcp
PII Filtering
Patient-identifying tags can be redacted from all tool output by setting an environment variable:
export DICOM_MCP_PII_FILTER=true
When enabled, the following tags are replaced with [REDACTED] in tool responses:
- PatientName
- PatientID
- PatientBirthDate
- PatientSex
This affects dicom_get_metadata, dicom_compare_headers, dicom_summarize_directory, and dicom_query. All other tools do not expose patient data and are unaffected.
To disable, unset the variable or set it to any value other than true, 1, or yes.
Project Structure
dicom_mcp/ # Python package
__init__.py # Re-exports all public symbols
__main__.py # Entry point: python -m dicom_mcp
server.py # FastMCP instance and run()
config.py # Environment-driven configuration
constants.py # Enums and tag group definitions
pii.py # PII redaction functions
helpers/ # Internal helper functions
tags.py # Tag reading, formatting, resolution
sequence.py # Sequence type identification
files.py # DICOM file discovery
philips.py # Philips private tag resolution
pixels.py # Pixel array and statistics
filters.py # Search filter parsing
tree.py # DICOM tree building and formatting
tools/ # MCP tool definitions
discovery.py # dicom_list_files, dicom_find_dixon_series
metadata.py # dicom_get_metadata, dicom_compare_headers
query.py # dicom_query, dicom_summarize_directory
validation.py # dicom_validate_sequence, dicom_analyze_series
search.py # dicom_search
philips.py # dicom_query_philips_private
pixels.py # dicom_read_pixels, dicom_compute_snr, dicom_render_image
tree.py # dicom_dump_tree
uid_comparison.py # dicom_compare_uids
segmentation.py # dicom_verify_segmentations
ti_analysis.py # dicom_analyze_ti
docs/ # Documentation
USAGE.md # Detailed tool reference and QA workflows
TODO.md # Planned improvements and known issues
CAPABILITIES.md # Plain-English summary of all capabilities
GUIDELINES.md # Behavioural constraints and regulatory context
dicom_mcp.py # Backward-compatible entry point shim
test_dicom_mcp.py # Test suite (138 tests)
Development
Running Tests
poetry run pytest -v --tb=short
The test suite includes 138 tests covering tool registration, helper functions, PII filtering, and full pipeline integration tests using synthetic DICOM files.
Linting
ruff check dicom_mcp/ dicom_mcp.py test_dicom_mcp.py
black --check dicom_mcp/ dicom_mcp.py test_dicom_mcp.py
Building Standalone Packages
# macOS (detects architecture automatically)
./build_standalone.sh
# Linux
./build_standalone_linux.sh
This bundles Python 3.12, fastmcp, pydicom, numpy, and Pillow into a self-contained package (~50MB) that requires no system Python.
MCP Inspector
To test the server interactively outside of Claude:
npx @modelcontextprotocol/inspector poetry run python -m dicom_mcp
Extending the Server
New tools can be added by creating a module in dicom_mcp/tools/ and importing it from dicom_mcp/tools/__init__.py. Each tool module imports the shared mcp instance from dicom_mcp.server and uses the @mcp.tool() decorator.
Other extension points:
- Custom tag groups: Edit
COMMON_TAGSindicom_mcp/constants.py - Sequence type detection: Modify
_identify_sequence_type()indicom_mcp/helpers/sequence.py - PII tag set: Edit
PII_TAGSindicom_mcp/pii.py(derived fromCOMMON_TAGS["patient_info"])
Troubleshooting
"Module not found" errors
Install all dependencies:
poetry install --with dev
If the MCP server starts but fails with ModuleNotFoundError for numpy or Pillow, the Claude Desktop config may be pointing to a different Python than the Poetry virtualenv. Either install the missing packages into that Python or update the config to use the virtualenv Python directly (see INSTALL.md).
"File not found" errors
Use absolute paths, not relative paths. Check that the path exists and is accessible.
"Not a valid DICOM file" errors
Verify the file is actually a DICOM file. Try opening with another DICOM viewer or pydicom.dcmread() directly.
Server not appearing in Claude Desktop
- Verify the configuration file path is correct
- Check that the configured Python can find all dependencies
- Restart Claude Desktop after configuration changes
- Check Claude Desktop logs for error messages
"No pixel data" errors
Some DICOM files (presentation states, structured reports, derived objects) don't contain pixel data. Use dicom_get_metadata to check the file's SOP Class or ImageType before attempting pixel operations.
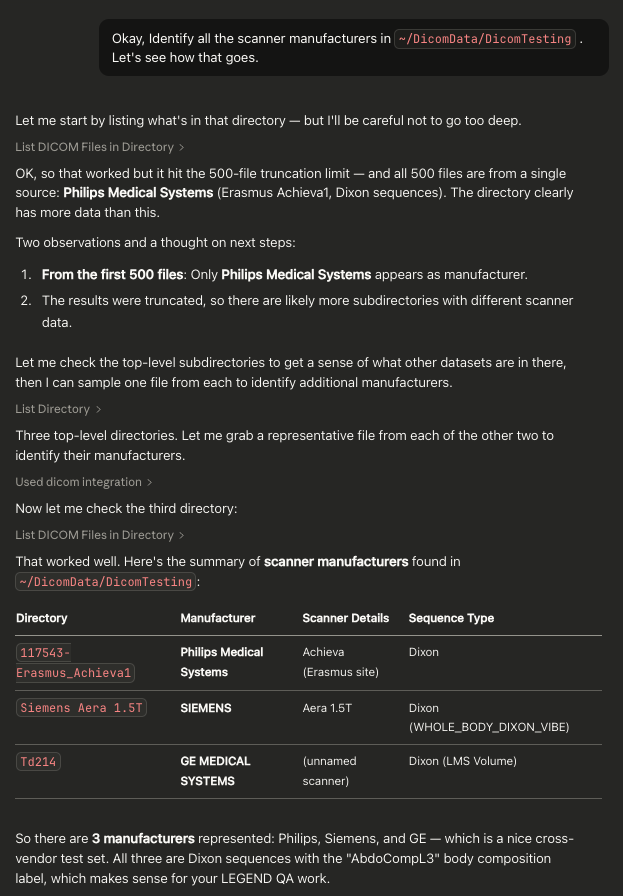
Example Screenshot
License
This MCP server is provided as-is for QA and testing purposes.
Support
For issues or questions:
- Check the Troubleshooting section above
- See docs/USAGE.md for detailed tool reference and workflow examples
- See INSTALL.md for installation help
- Verify your DICOM files with pydicom directly
- Review MCP server logs in Claude Desktop