24 KiB
DICOM MCP Server — Usage Guide
Detailed usage examples, tool reference, and QA workflow patterns for the DICOM MCP server.
For installation and setup, see INSTALL.md. For project overview, see README.md.
Tip: Pair with a filesystem MCP server for maximum capability. The DICOM MCP server works well on its own for metadata inspection and validation. However, if Claude also has access to a filesystem MCP server with media reading support (e.g. the
read_media_filetool), it can view rendered images directly, enabling an interactive visual feedback loop — render an image, inspect it, adjust ROI placement, measure, iterate — all within a single conversation. Without a filesystem MCP, Claude can still render images and save them to disk, but you'll need to open them yourself in a viewer and describe what you see.
Quick Examples
Finding test data
List all Dixon sequences in my test data directory
The server will use dicom_find_dixon_series to locate all Dixon sequences and show what image types are available.
Comparing fat/water selection
Compare the headers of these two files to see which Dixon images were selected:
- /data/test01/water.dcm
- /data/test01/fat.dcm
Validating a protocol
Validate that /data/scan01/image001.dcm matches our Siemens protocol:
- TR should be 4.5
- TE should be 2.3
- Manufacturer should be Siemens
Quick directory overview
Summarize what's in the /data/large_study directory
Finding specific files
Find all files in /data/study where the series description contains "MOST"
and echo time is greater than 10
Rendering an image
Render /data/liver_scan/slice_005.dcm with auto-windowing
Measuring SNR
Compute the SNR on /data/scan.dcm with a signal ROI at [100, 200, 50, 50]
in the liver and a noise ROI at [20, 20, 40, 40] in background air
Dumping full DICOM structure
Dump the full DICOM tree of /data/scan.dcm including nested sequences
Comparing UIDs across directories
Compare the SeriesInstanceUIDs between /data/study_v1 and /data/study_v2
Verifying segmentation references
Check that all segmentation files in /data/segs reference valid source images
Analysing MOLLI inversion times
Analyze the inversion times in /data/molli_series — is it a proper 5(3)3 scheme?
Inspecting Philips private tags
List all Philips Private Creator tags in /data/philips_scan/image001.dcm
Look up DD 001 element offset 0x85 in /data/philips_scan/image001.dcm
PII Filtering
When PII filtering is enabled (via DICOM_MCP_PII_FILTER=true), the following patient tags are replaced with [REDACTED] in tool output:
- PatientName
- PatientID
- PatientBirthDate
- PatientSex
Affected tools: dicom_get_metadata, dicom_compare_headers, dicom_summarize_directory, dicom_query.
All other tools do not expose patient data and are unaffected. See INSTALL.md for configuration instructions.
Tool Reference
Directory & File Discovery
dicom_list_files
List all DICOM files in a directory with metadata filtering.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path to search |
recursive |
bool | true |
Search subdirectories |
filter_sequence_type |
string | null |
Filter by type: dixon, t1_mapping, multi_echo_gre, spin_echo_ir, t1, t2, flair, dwi, localizer |
count_only |
bool | false |
Return series breakdown with file counts instead of individual file listing |
response_format |
string | markdown |
markdown or json |
{
"directory": "/data/test_suite",
"filter_sequence_type": "dixon",
"count_only": true
}
dicom_summarize_directory
Get a high-level overview of DICOM directory contents showing unique values and file counts for patient info, manufacturer, scanner model, field strength, institution, series descriptions, and sequence types.
When PII filtering is enabled, patient tags (Name, ID, DOB, Sex) are redacted.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path to directory |
recursive |
bool | true |
Search subdirectories |
include_series_overview |
bool | true |
Include per-series table with acquisition parameters (TR, TE, TI, FA) |
response_format |
string | markdown |
markdown or json |
{
"directory": "/data/large_study",
"include_series_overview": true
}
dicom_find_dixon_series
Find and analyse Dixon (chemical shift) sequences. Automatically identifies Dixon series and detects image types (water, fat, in-phase, out-phase).
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path to search |
recursive |
bool | true |
Search subdirectories |
response_format |
string | markdown |
markdown or json |
{
"directory": "/data/body_comp_study"
}
dicom_search
Find DICOM files matching specific filter criteria using AND logic across multiple filters.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path to directory |
filters |
list[string] | required | Filter expressions (see syntax below) |
recursive |
bool | true |
Search subdirectories |
mode |
string | summary |
count, paths, or summary |
response_format |
string | markdown |
markdown or json |
Filter syntax:
| Operator type | Examples |
|---|---|
| Text (case-insensitive) | "SeriesDescription contains MOST", "PatientName is not UNKNOWN", "SeriesDescription starts with Thigh", "SeriesDescription ends with Dixon" |
| Numeric/symbolic | "EchoTime > 10", "FlipAngle <= 15", "RepetitionTime = 516", "SeriesNumber != 0" |
| Presence | "InversionTime exists", "InversionTime missing" |
Tags can be keywords (EchoTime) or hex codes (0018,0081).
{
"directory": "/data/study",
"filters": ["SeriesDescription contains MOST", "EchoTime > 10"],
"mode": "paths"
}
Metadata & Validation
dicom_get_metadata
Extract DICOM header information organised by tag groups. When PII filtering is enabled, patient tags are redacted.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to DICOM file |
tag_groups |
list[string] | null |
Groups to extract: patient_info, study_info, series_info, image_info, acquisition, manufacturer, equipment, geometry, pixel_data |
custom_tags |
list[string] | null |
Additional tags as hex codes, e.g. ["0018,0080"] |
philips_private_tags |
list[dict] | null |
Philips private tags to resolve (see below) |
response_format |
string | markdown |
markdown or json |
Philips private tags:
For Philips DICOM files, you can resolve private tags inline without using the separate dicom_query_philips_private tool. Each entry is a dict with dd_number and element_offset (and optionally private_group, default 0x2005).
{
"file_path": "/data/philips_scan.dcm",
"tag_groups": ["acquisition", "manufacturer"],
"philips_private_tags": [
{"dd_number": 1, "element_offset": 133}
]
}
Standard example:
{
"file_path": "/data/scan.dcm",
"tag_groups": ["acquisition", "manufacturer"],
"custom_tags": ["0018,0080", "0018,0081"]
}
dicom_compare_headers
Compare DICOM headers across multiple files side-by-side. When PII filtering is enabled, patient tags are redacted.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_paths |
list[string] | required | 2-10 files to compare |
tag_groups |
list[string] | ["acquisition", "series_info"] |
Which tag groups to compare |
show_differences_only |
bool | false |
Only show tags that differ |
response_format |
string | markdown |
markdown or json |
{
"file_paths": [
"/data/test01/water.dcm",
"/data/test01/fat.dcm",
"/data/test01/in_phase.dcm"
],
"show_differences_only": true
}
dicom_validate_sequence
Validate MRI sequence parameters against expected values.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | DICOM file to validate |
expected_parameters |
dict | null |
Expected values. Supported keys: RepetitionTime, EchoTime, InversionTime, FlipAngle, ScanningSequence, SequenceVariant, MRAcquisitionType |
manufacturer |
string | null |
Expected manufacturer name |
response_format |
string | markdown |
markdown or json |
{
"file_path": "/data/test.dcm",
"expected_parameters": {
"RepetitionTime": 4.5,
"EchoTime": 2.3,
"InversionTime": 100,
"FlipAngle": 10
},
"manufacturer": "Siemens"
}
dicom_analyze_series
Comprehensive analysis of a complete DICOM series — checks parameter consistency across all files, verifies completeness (no missing instances), and identifies anomalies.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path containing series files |
response_format |
string | markdown |
markdown or json |
{
"directory": "/data/series_001"
}
dicom_query
Query arbitrary DICOM tags across all files in a directory. Aggregates unique values with file counts. When PII filtering is enabled, patient tags are redacted.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Path to directory |
tags |
list[string] | required | Tag keywords (e.g. "EchoTime") or hex codes (e.g. "0018,0081") |
recursive |
bool | true |
Search subdirectories |
group_by |
string | null |
Optional tag to group results by (e.g. "SeriesDescription") |
response_format |
string | markdown |
markdown or json |
{
"directory": "/data/study",
"tags": ["EchoTime", "FlipAngle"],
"group_by": "SeriesDescription"
}
Structure & Comparison
dicom_dump_tree
Full hierarchical dump of DICOM structure, including nested sequences (SQ elements) with tree-character formatting. Useful for understanding complex DICOM files, inspecting nested structures (e.g. ReferencedSeriesSequence, SourceImageSequence), and debugging.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to DICOM file |
max_depth |
int | 10 |
Maximum nesting depth to display |
show_private |
bool | true |
Include private tags in output |
response_format |
string | markdown |
markdown or json |
When PII filtering is enabled, patient tags are redacted in the tree output. Pixel data (7FE0,0010) is always skipped.
{
"file_path": "/data/scan.dcm",
"max_depth": 5,
"show_private": false
}
dicom_compare_uids
Compare UID sets between two DICOM directories. Identifies shared UIDs, UIDs unique to each directory, and reports counts and details.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory1 |
string | required | First directory to compare |
directory2 |
string | required | Second directory to compare |
recursive |
bool | true |
Search subdirectories |
compare_tag |
string | SeriesInstanceUID |
Tag to compare — keyword (e.g. StudyInstanceUID, SOPInstanceUID) or hex code (e.g. 0020,000E) |
response_format |
string | markdown |
markdown or json |
{
"directory1": "/data/study_original",
"directory2": "/data/study_reprocessed",
"compare_tag": "SOPInstanceUID"
}
Segmentation & T1 Mapping
dicom_verify_segmentations
Validate that segmentation files correctly reference valid source images. Detects segmentation files by SOPClassUID (1.2.840.10008.5.1.4.1.1.66.4) or by the presence of SourceImageSequence. For each segmentation, checks that every ReferencedSOPInstanceUID points to an existing source file in the same directory.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Directory containing segmentation and source files |
recursive |
bool | true |
Search subdirectories |
response_format |
string | markdown |
markdown or json |
Reports total segmentation files, total references, matched vs unmatched counts, and details for any unmatched references.
{
"directory": "/data/study_with_segmentations"
}
dicom_analyze_ti
Extract and validate inversion times from MOLLI / T1 mapping sequences. Handles vendor-specific differences automatically:
- Siemens / GE: reads the standard
InversionTimetag(0018,0082) - Philips: falls back to private tag DD 006, offset 0x72 in group 2005 (confirmed at
(2005,xx72)) or DD 001, offset 0x20 in group 2001
Groups by series, sorts by instance number, and computes statistics including gap analysis to detect missing or out-of-range inversion times.
| Parameter | Type | Default | Description |
|---|---|---|---|
directory |
string | required | Directory containing MOLLI / T1 mapping files |
recursive |
bool | true |
Search subdirectories |
gap_threshold |
float | 2500.0 |
Flag consecutive TI gaps exceeding this value (ms) |
response_format |
string | markdown |
markdown or json |
Output includes per series:
- Ordered TI list sorted by instance number
- Count of zero-value TIs (typically output maps, not acquisitions)
- Count of non-zero TIs (actual inversions)
- TI range, mean, and median
- Largest consecutive gap, with warning if exceeding the threshold
{
"directory": "/data/molli_series",
"gap_threshold": 3000.0
}
Philips Private Tags
dicom_query_philips_private
Query Philips private DICOM tags using DD number and element offset. Philips MRI scanners store proprietary metadata in private tag blocks whose assignments vary across scanners and software versions — this tool resolves them dynamically.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to a Philips DICOM file |
dd_number |
int | null |
DD number to look up (e.g. 1 for "DD 001") |
element_offset |
int | null |
Element offset within the DD block (e.g. 133 for 0x85) |
private_group |
int | 0x2005 |
DICOM private group to search |
list_creators |
bool | false |
List all Private Creator tags instead of resolving a specific one |
response_format |
string | markdown |
markdown or json |
Two usage modes:
- List creators — discover what private tag blocks are available:
{
"file_path": "/data/philips_scan.dcm",
"list_creators": true
}
- Resolve a specific tag — look up a known DD number and offset:
{
"file_path": "/data/philips_scan.dcm",
"dd_number": 1,
"element_offset": 133
}
Common Philips DD numbers and offsets (group 2005):
| DD # | Offset | Description |
|---|---|---|
| 001 | 0x85 | Shim calculation values |
| 001 | 0x63 | Stack ID |
| 004 | 0x00 | MR Series data object |
Pixel Analysis
dicom_read_pixels
Extract pixel statistics from a DICOM file. Values are rescaled using RescaleSlope and RescaleIntercept when present (standard for Philips, common on Siemens/GE).
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to DICOM file |
roi |
list[int] | null |
Region of interest as [x, y, width, height] (top-left corner in pixel coordinates). If omitted, statistics cover the entire image. |
include_histogram |
bool | false |
Include a binned histogram of pixel values |
histogram_bins |
int | 50 |
Number of histogram bins |
response_format |
string | markdown |
markdown or json |
Returns: min, max, mean, standard deviation, median, 5th/25th/75th/95th percentiles, and pixel count.
{
"file_path": "/data/liver_scan/slice_005.dcm",
"roi": [100, 200, 50, 50],
"include_histogram": true,
"histogram_bins": 20
}
dicom_compute_snr
Compute signal-to-noise ratio from two ROIs in a DICOM image using the single-image method: SNR = mean(signal) / std(noise).
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to DICOM file |
signal_roi |
list[int] | required | Signal region as [x, y, width, height] |
noise_roi |
list[int] | required | Noise/background region as [x, y, width, height] |
response_format |
string | markdown |
markdown or json |
Tip: Use dicom_render_image with overlay_rois first to visualise and verify ROI placement before measuring.
Note: Some manufacturers (notably Philips) zero-fill background air outside the reconstruction FOV, which results in zero noise standard deviation and infinite SNR. In such cases, consider using a homogeneous tissue region (e.g. subcutaneous fat or muscle) as the noise ROI instead.
{
"file_path": "/data/liver_scan/slice_005.dcm",
"signal_roi": [100, 200, 50, 50],
"noise_roi": [20, 20, 40, 40]
}
dicom_render_image
Render a DICOM image to PNG with configurable windowing. Optionally overlays labelled ROI rectangles for visual verification.
| Parameter | Type | Default | Description |
|---|---|---|---|
file_path |
string | required | Path to DICOM file |
output_path |
string | required | Path where the PNG will be saved |
window_center |
float | null |
Manual window center |
window_width |
float | null |
Manual window width |
auto_window |
bool | false |
Auto-calculate window from 5th-95th percentile |
overlay_rois |
list[dict] | null |
ROI overlays (see below) |
show_info |
bool | true |
Burn in series description and windowing values |
response_format |
string | markdown |
markdown or json |
Windowing priority:
- Explicit
window_center+window_widthparameters auto_window=True→ 5th-95th percentile range- DICOM header WindowCenter/WindowWidth values
- Fallback → full pixel range (min to max)
ROI overlay format:
{
"roi": [x, y, width, height],
"label": "Signal (Liver)",
"color": "green"
}
Available colours: red, green, blue, yellow, cyan, magenta, white.
Full example:
{
"file_path": "/data/scan.dcm",
"output_path": "/data/renders/scan_auto.png",
"auto_window": true,
"overlay_rois": [
{"roi": [100, 200, 50, 50], "label": "Signal", "color": "green"},
{"roi": [20, 20, 40, 40], "label": "Noise", "color": "red"}
]
}
Viewing rendered images: If Claude also has access to a filesystem MCP server with media reading capability (e.g. read_media_file), it can view the rendered PNG directly and use visual feedback to refine ROI placement iteratively — no need to switch to a separate DICOM viewer.
Output Formats
All tools support two output formats via the response_format parameter:
Markdown (default) — Human-readable with tables, formatting, and visual indicators. Best for conversational use in Claude Desktop.
JSON — Machine-readable structured data with consistent schemas. Best for programmatic processing or piping into other tools.
Performance Considerations
File scanning limits
All directory scanning tools are limited to processing 1000 files per scan by default. This limit is configurable via the DICOM_MCP_MAX_FILES environment variable (see INSTALL.md). If this limit is reached, a warning indicates results were truncated (and truncated: true in JSON output). Narrow your search to a more specific subdirectory if needed.
Optimisation tips
- Use
dicom_summarize_directoryinstead ofdicom_list_fileswhen you only need an overview - Use
dicom_list_fileswithcount_only: truefor a compact series inventory - Use
dicom_searchwithmode: "count"to quickly check match counts before fetching details - Use
dicom_queryinstead ofdicom_get_metadataon multiple files when you need aggregate tag values - Set
recursive: falseto scan only the immediate directory - For pixel tools, use
dicom_render_imageto verify ROI placement before runningdicom_compute_snr - Use
dicom_dump_treewithshow_private: falseto focus on standard DICOM structure - Use
dicom_compare_uidsto quickly detect differences between study directories without inspecting every file
QA Workflow Examples
Workflow 1: Dixon Image Selection Investigation
When you suspect incorrect fat/water selection:
dicom_find_dixon_series— locate the Dixon seriesdicom_list_fileswithfilter_sequence_type: "dixon"— see all Dixon filesdicom_compare_headerson suspected files — focus on ImageType tagsdicom_get_metadata— extract full headers for documentation
Workflow 2: Multi-Manufacturer Validation
When testing across GE, Siemens, and Philips:
dicom_summarize_directory— check patient info consistency and get a quick inventorydicom_querywithgroup_by: "SeriesDescription"— compare timing parameters across seriesdicom_validate_sequencewith manufacturer-specific expected parametersdicom_compare_headers— identify parameter variations- Document differences for test specification updates
Workflow 3: Test Dataset Verification
Before running automated tests:
dicom_analyze_serieson each test case directorydicom_find_dixon_seriesto confirm Dixon sequences are presentdicom_validate_sequenceto check protocol compliancedicom_searchto find files with unexpected parameter values (e.g."FlipAngle != 15")
Workflow 4: Image Quality Assessment
For checking signal quality and image rendering:
dicom_render_imagewithauto_window: true— render the image and visually inspectdicom_render_imagewithoverlay_rois— place and verify signal/noise ROI positionsdicom_read_pixels— check pixel value distributions and histogramsdicom_compute_snr— measure SNR with verified ROI placements- Repeat across series or manufacturers to compare image quality
Note on Philips data: Background air is often zero-filled, making the traditional background-air SNR method return infinite. Use a homogeneous tissue region (subcutaneous fat, muscle) as the noise proxy instead.
Workflow 5: MOLLI / T1 Mapping Validation
When verifying T1 mapping acquisitions across vendors:
dicom_analyze_ti— extract all inversion times, check for proper MOLLI scheme (e.g. 5(3)3 = 11 TI-weighted images + output maps)- Check for gap warnings — large gaps between consecutive TIs may indicate missing heartbeats or acquisition failures
dicom_dump_treeon a representative file — inspect nested sequence structure if TI extraction fails- For Philips data:
dicom_query_philips_privatewithdd_number: 6, element_offset: 114(0x72) — manually verify the private TI tag if needed
Workflow 6: Segmentation Verification
When validating segmentation files reference the correct source images:
dicom_verify_segmentations— check all segmentation-to-source references in one pass- Review any unmatched references — missing source files or incorrect ReferencedSOPInstanceUIDs
dicom_compare_uidswithcompare_tag: "SOPInstanceUID"— compare two directories to find missing or extra files
Workflow 7: Philips Private Tag Investigation
When investigating Philips-specific metadata:
dicom_query_philips_privatewithlist_creators: true— discover available DD blocksdicom_query_philips_privatewith specificdd_numberandelement_offset— resolve individual tags- Or use
dicom_get_metadatawith thephilips_private_tagsparameter to extract private tags alongside standard metadata in a single call
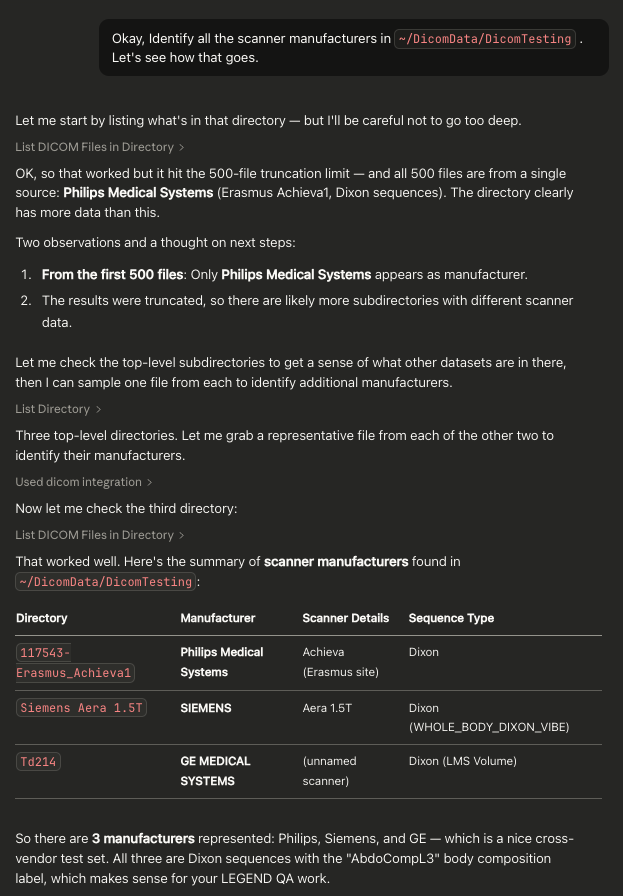
Example Screenshot From Claude Desktop